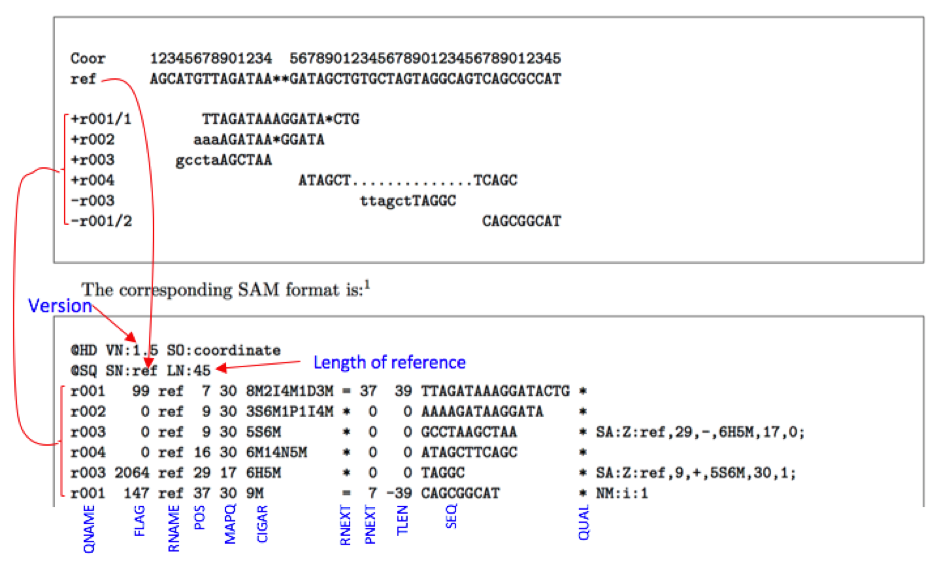

Header lines start with ‘’, while alignment lines do not. SAM is a TAB-delimited text format consisting of a header section and an alignment section. Keep in mind that this may take a half hour or more depending on the size of your bam and the speed of your computer. The SAM (Sequence Alignment/Map) format (BAM is just the binary form of SAM) is. A BAM file (.bam) is a compressed binary version (BGZF format) of a SAM file that is used to represent aligned sequences. If you have any doubt, though, it's easy enough to delete your bai file, then generate a new index using the previous command. If your index file is named identically, with just the additional ".bai" suffix, you can be reasonably sure that it was generated from the same file. If you have a bam file without a corresponding index, you can generate one using "samtools index bamfile.bam". Without the corresponding bam file, your bai file is useless, since it doesn't actually contain any sequence data. This file acts like an external table of contents, and allows programs to jump directly to specific parts of the bam file without reading through all of the sequences. While we will go into some features of the SAM format, the paper by Heng Li et al provides a lot more detail on the. This file has the same name, suffixed with. The output we requested from the Bowtie2 aligner is an unsorted SAM file, also known as Sequence Alignment Map format.The SAM file, is a tab-delimited text file that contains information for each individual read and its alignment to the genome. ecSeq is a bioinformatics solution provider with solid expertise in the analysis of high-throughput sequencing data. You can view what's in the bam file using "samtools view bamfile.bam | less".īam files can also have a companion file, called an index file. A bai file isn't an indexed form of a bam - it's a companion to your bam that contains the index.Ī bam file is a binary blob that stores all of your aligned sequence data.

